Telomere Analysis
Telomeres are fundamental functional elements of eukaryotic chromosomes involved in the maintenance of genome stability, and their length affects many cellular processes, such as senescence, carcinogenesis, and senescence. Telomere length is maintained through a dynamic process of shortening and lengthening, and significantly shortened chromosome ends can lead to telomere-initiated senescence or apoptosis. Loss of telomere length and function may be due to normal shortening of telomeres (in cells lacking telomerase), loss of protective function, or altered DNA repair. The most important indicator of correct telomere function is the maintenance of telomere length within the range typical for each species. The classic method of telomere length assessment is through the classical method based on Southern blotting, although this method is relatively crude and can only provide an estimate of average telomere length. To overcome the shortcomings of classical telomere length estimation based on Southern blotting, Q-FISH was invented. Quantitative FISH (Q-FISH) is a comprehensive method that measures individual chromosome telomere length in individual cells with a resolution of 200 base pairs. State-of-the-art methods use peptide nucleic acid (PNA) telomeric oligonucleotide probes and appropriate digital imaging software to capture and quantify fluorescent signals.
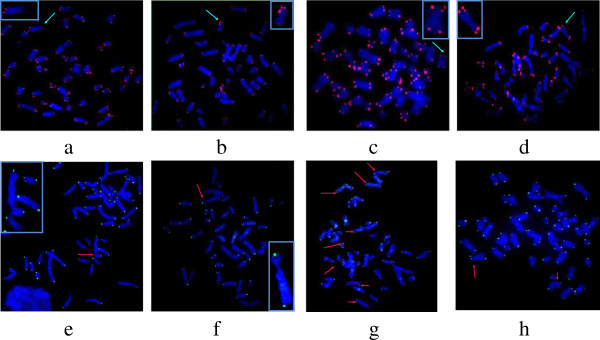 Fig 1.Representative images of a metaphase spread hybridized with telomeric PNA probes using Q-FISH and CO-FISH protocols. (Joksic I, et al. 2012)
Fig 1.Representative images of a metaphase spread hybridized with telomeric PNA probes using Q-FISH and CO-FISH protocols. (Joksic I, et al. 2012)
Q-FISH Services
Telomere length is variable between species and even heterogeneous within different cell types of an individual. Chromosomes require a certain number of telomeric (TTAGGG)n repeats to “cover” the chromosome and avoid activating the DNA damage response. Uncapped telomeres are considered double-strand breaks, which lead to the activation of two major DNA damage response mechanisms (homologous recombination (HR) and non-homologous end-joining (NHEJ)). The basic workflow of our Q-FISH testing service involves immobilizing the target metaphase chromosomes to the surface of a slide, denaturing them, hybridizing them with probes, and then imaging them by washing and counterstaining. Analyzed using image capture software and telomere measurement tools, this protocol contains an internal control of fluorescence intensity for improved accuracy. The accuracy of the Q-FISH method is corrected for the fluorescence intensity of the telomere signal by the fluorescence intensity of the internal control using sufficient internal controls to avoid imprecision due to changes in lamp intensity. Additional also allows conversion of fluorescence intensity values to DNA length units (RTLU) by telomere/centromere ratios or calibration relative to cultured cells of known telomere length. Our service platform can provide the above two calibration methods. Through this service, researchers can study molecular mechanisms related to telomere length regulation, cell cycle, cell lifespan, and more.
 Fig 2. Telomere length assessment by Q-FISH access.
Fig 2. Telomere length assessment by Q-FISH access.
Applications
- Q-FISH can be used to determine telomere length (p and q arms) in a given cell;
- Q-FISH can be used to determine the intrachromosomal distribution of the shortest/longest telomeres;
- Q-FISH can be used to estimate telomere length in species containing interstitial telomeric sites in the genome;
- Q-FISH can be used to estimate telomere length in species with ultralong and heterogeneous telomeres, such as mice;
Creative Bioarray provides Q-FISH-based telomere length assessment services. You will find solutions to your research needs through our technology platform. If you are interested in our service, please contact us for cooperation. We look forward to cooperating with you in the near future.
Reference
- Joksic I, Vujic D, Guc-Scekic M, et al. Dysfunctional telomeres in primary cells from Fanconi anemia FANCD2 patients[J]. Genome integrity, 2012, 3(1): 1-12.
All products and services on this website are only suitable for non-medical purposes.


 Fig 1.Representative images of a metaphase spread hybridized with telomeric PNA probes using Q-FISH and CO-FISH protocols. (Joksic I, et al. 2012)
Fig 1.Representative images of a metaphase spread hybridized with telomeric PNA probes using Q-FISH and CO-FISH protocols. (Joksic I, et al. 2012) Fig 2. Telomere length assessment by Q-FISH access.
Fig 2. Telomere length assessment by Q-FISH access.


