sRNA Research and FISH
sRNA Research
An important information link in the process of RNA gene expression, and the location of RNA information is essential for understanding its function. The understanding of gene function has gradually expanded the horizon, from focusing only on coding sequences to the study of non-coding RNA (ncRNA). Especially after discovering that ncRNA is essential for the regulation of gene expression. Generally, ncRNA is simply divided into small RNA (sRNA) and long non-coding RNA (lncRNA) in terms of sequence length. The length of sRNA is 18 to 30 nt and usually contains 18–24 nt microRNA (miRNA), 21-22 nt short interfering RNA (siRNA) and 26–30 nt PIWI-associated RNA (piRNA). Their small size and high diversity make it challenging to develop detection methods with sufficient resolution and specificity for multiplexing and quantification. The subcellular localization of the components of the miRNA and siRNA pathway has been described, including Cajal bodies (CB), cleavage bodies, processing bodies (P bodies), and the recently described membrane-bound polysomes. sRNA can function in a homology-dependent manner to direct transcription and post-transcriptional silencing.
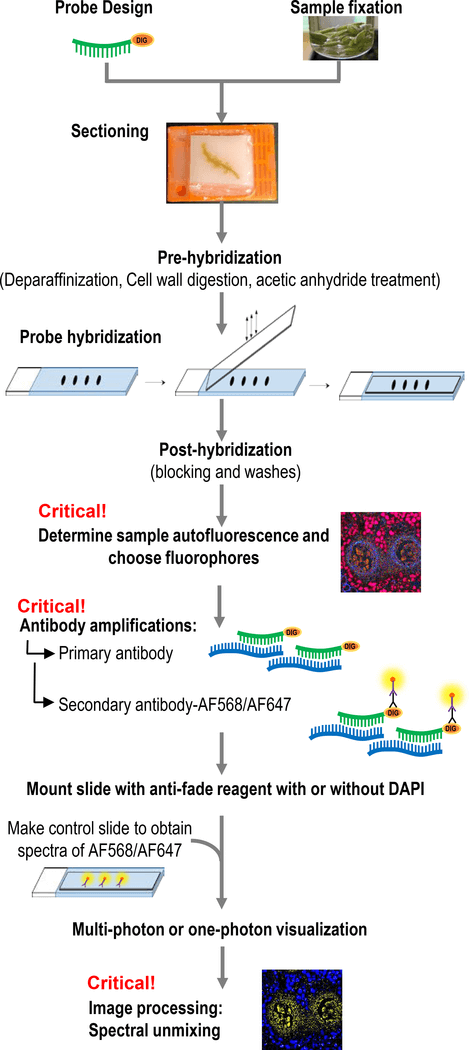 Fig 1. Workflow of sRNA-FISH. (Huang K, et al. 2019)
Fig 1. Workflow of sRNA-FISH. (Huang K, et al. 2019)
sRNA In Situ Visualization Analysis
RNA-FISH is a powerful method for detecting specific RNA, allowing the localization and quantification of RNA molecules within individual cells and tissues. We provide comprehensive sRNA analysis services for samples from plants, animals, and microorganisms. Studies have shown that plants and animals have their own ways to produce developmentally and functionally important sRNAs. The formation of sRNA in animal cells mainly occurs in the nucleus and cytoplasm. In plant cells, miRNA processing occurs in the nucleus in most cases. Studies have shown that plants and animals have their own ways to produce developmentally and functionally important small RNAs (sRNAs). In animal cells, cleavage of siRNA and miRNA occurs in the nucleus and cytoplasm. In plant cells, miRNA processing occurs in the nucleus in most cases. sRNA can function in a homology-dependent manner to guide transcription and post-transcriptional silencing and to regulate the efficiency of gene expression.
 Fig 2. Flow chart of the RNA-FISH.
Fig 2. Flow chart of the RNA-FISH.
sRNA Visualization Analysis Strategy
In the analysis of sRNA, researchers have begun to use advanced locked nucleic acid (LNA) probes. LNAs are a new class of RNA analogs with a high affinity for complementary DNA and RNA. LNA has several advantages over DNA oligonucleotide probes, for example, it has been shown to significantly increase the stability of its thermal duplexes and improve the distinction between perfectly matched and mismatched target nucleic acids. These characteristics help to cope with the characteristics of "small scale, high diversity".
| Service Sub-items | Key Point | Technology Features |
| miRNA-FISH Service | miRNA+mRNA | - Advanced LNA/DNA probes for FISH detection.
- Use a combination of antibodies and probes conjugated to different fluorophores to detect two sRNAs in the same sample.
- It can not only realize single target detection but also detect more than 2 targets at the same time.
|
| siRNA-FISH Service | siRNA+mRNA |
Creative Bioarray provides a wide range of RNA-FISH analysis services, from program customization to graphic processing, a variety of technical service options to help our customers improve research efficiency. If you are interested in our FISH analysis services, please contact us for cooperation. We look forward to cooperating with you in the near future.
References
- Huang K, Baldrich P, Meyers B C, et al. sRNA‐FISH: versatile fluorescent in situ detection of small RNAs in plants[J]. The Plant Journal, 2019, 98(2): 359-369.
- Huang K, Demirci F, Batish M, et al. Quantitative, super-resolution localization of small RNAs with sRNA-PAINT[J]. Nucleic acids research, 2020, 48(16): e96-e96.
All products and services on this website are only suitable for non-medical purposes.


 Fig 1. Workflow of sRNA-FISH. (Huang K, et al. 2019)
Fig 1. Workflow of sRNA-FISH. (Huang K, et al. 2019) Fig 2. Flow chart of the RNA-FISH.
Fig 2. Flow chart of the RNA-FISH.


