DNA-RNA FISH Analysis
Studying how RNA interacts with its DNA target in the cellular environment is increasingly important for understanding transcription events. Many studies have shown that the spatial location of genes in the interphase nucleus is highly related to their functions. Fluorescence in situ hybridization (FISH) is a powerful molecular technique that can be used to map gene loci in the nucleus of interphase cells to detect nucleic acids in cells. The FISH method can detect DNA or RNA, but both methods have certain limitations. DNA FISH can only detect the physical location of genes, not gene activity; while RNA FISH detects transcripts, but may miss a large number of alleles, because not all alleles of a gene must be transcribed at the same time. The most effective method for mapping gene loci and their activity is sequence RNA and DNA FISH. This is an important technique to reveal how gene mapping is related to the activity. Combining DNA and RNA FISH in the same cell is technically challenging because the conditions suitable for DNA FISH may be too harsh for fragile single-stranded RNA molecules.
Sequential DNA-RNA FISH Services
Simultaneous DNA and RNA analysis have been used in a variety of DNA and transcription analysis experiments. For example, the accumulation of nascent transcripts can be visualized near activated genomic sites by RNA/DNA fluorescence in situ hybridization (FISH). Due to the unique properties of DNA and RNA, the use of sequential RNA-DNA FISH to quantify the changes in RNA transcription between alleles in a single cell nucleus requires a customized protocol when performing a single FISH analysis. The process of DNA FISH may damage the level of RNA FISH signal, so we use tyramide signal amplification (TSA) to avoid the loss of RNA signal in the DNA/RNA FISH combination scheme. In addition, it can also be solved by sequential FISH.
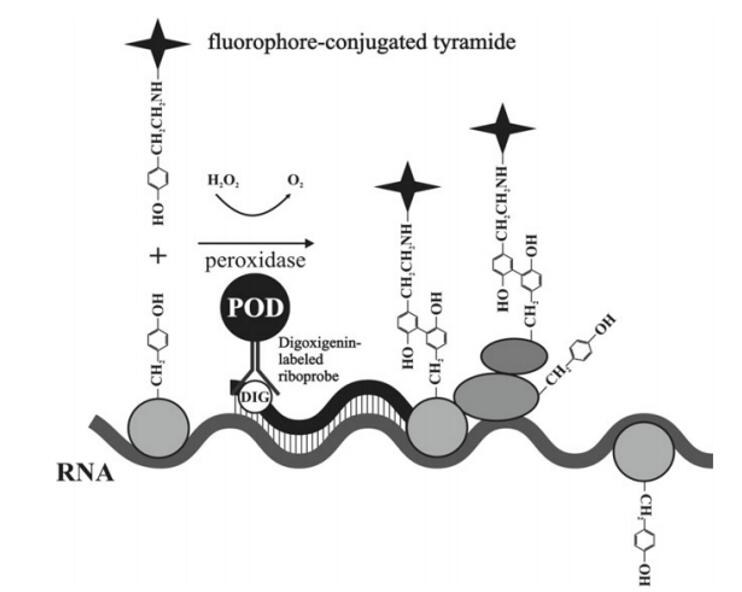 Fig 1. Mechanism of tyramide signal amplifi cation (TSA). (Shpiz S, et al. 2014)
Fig 1. Mechanism of tyramide signal amplifi cation (TSA). (Shpiz S, et al. 2014)
Services
Our DNA-RNA FISH service is an analysis service designed for each customer's samples and experiments. The service process includes experimental plan design, probe customization, sample pre-processing, FISH hybridization, microscope services and data analysis services.
 Fig 2. Our DNA-RNA FISH service.
Fig 2. Our DNA-RNA FISH service.
Option 1: Sequential FISH
This strategy can first detect RNA and then perform DNA detection. We use Tyramine Signal Amplification (TSA) to detect RNA FISH signals in the combined DNA/RNA FISH protocol to protect the RNA FISH signals in subsequent DNA FISH procedures. First, use a biotin-labeled single-stranded DNA probe for RNA detection, and use TSA to visualize the hybridization probe. After the signal is collected, it is processed by RNase and DNA FISH is performed. The signal enhancement of TSA may create some background in the nucleus and remaining cytoplasm, but DNA FISH distinguishes them from the active site.
Option 2: Simultaneous FISH
This service can be carried out using mixed probe hybridization or the introduction of immunostaining. The advantage of using mixed FISH probes for hybridization analysis is that it can shorten the test cycle and is suitable for some fragile samples that are not suitable for undergoing a large number of processing steps. The method of introducing immunostaining is based on the amino-labeled oligonucleotide probes that are first hybridized with RNA probes and then need to be fixed and denatured and then hybridized with DNA probes.
 Fig 3. Features of our DNA-RNA FISH service.
Fig 3. Features of our DNA-RNA FISH service.
Creative Bioarray provides one-stop DNA-RNA FISH technical services to help our customers speed up turnover. You will benefit from our technical expertise and experimental platform, just mail the sample to us and confirm the plan given by our laboratory. We will provide you with icons and raw data files for publication at the end of the analysis. If you are interested in our DNA-RNA FISH service, please contact us for cooperation. We look forward to cooperating with you in the near future.
References
- Takizawa T, Misteli T. Sequential RNA and DNA fluorescence in situ hybridization (PROT39)[J].
- Shpiz S, Lavrov S, Kalmykova A. Combined RNA/DNA fluorescence in situ hybridization on whole-mount Drosophila ovaries[M]//PIWI-Interacting RNAs. Humana Press, Totowa, NJ, 2014: 161-169.
- Magaraki A, Loda A, Gribnau J, et al. Simultaneous RNA–DNA FISH in Mouse Preimplantation Embryos[M]//X-Chromosome Inactivation. Humana, New York, NY, 2018: 131-147.


 Fig 1. Mechanism of tyramide signal amplifi cation (TSA). (Shpiz S, et al. 2014)
Fig 1. Mechanism of tyramide signal amplifi cation (TSA). (Shpiz S, et al. 2014) Fig 2. Our DNA-RNA FISH service.
Fig 2. Our DNA-RNA FISH service. Fig 3. Features of our DNA-RNA FISH service.
Fig 3. Features of our DNA-RNA FISH service.


