Creative Bioarray provides a comprehensive visual analysis solution to help customers solve the ambiguity of breast cancer-related HER2 expression and further analysis of heterogeneous samples. This service can be used to verify breast cancer-related transcriptome data and analyze the subcellular localization of various RNAs in breast cancer (including various ncRNAs such as mRNA and lncRNA). In addition, services can also be carried out at the verification stage of preclinical drug efficacy evaluation and evaluation of potential prognostic markers for breast cancer treatment.
FISH Analysis in Breast Cancer
The diagnosis and characterization of breast cancer involves evaluating the biomarkers found in or on tumor cells, the three most important of which are estrogen receptor (ER), progesterone receptor (PR), and human epidermis Growth factor receptor 2 (ERBB2, formerly known as HER2). The amplification of the ERBB2 gene located on chromosome 17 (17q12) or the overexpression of this receptor has an important impact on the development of invasive breast cancer. It is important to detect human epidermal growth factor receptor 2 gene amplification in breast and other cancers. The copy number of the EGFR gene and mutations in EGFR or other downstream genes assessed by FISH have been studied as potential biomarkers that may predict treatment sensitivity.
According to the updated HER2 interpretation guide (ASCO-CAP), the ambiguous sample of HER2 is recommended for further analysis and testing. The HER2 amplicon at 17q12 contains multiple genes, which may be co-amplified with HER2. The HER2 amplicon at 17q12 contains multiple genes, which may be co-amplified with HER2. The use of alternative chromosome 17 probes allows another way to calculate the HER2:Chromosome 17 ratio, providing an alternative FISH ratio score in the case of complex CEP17 patterns. Alternative probes used in this part of the study include RAI1 (retinoic acid induction 1; formerly known as SMS), RARA (retinoic acid receptor), and TP53 (tumor protein 53). This service aims to help the research on the mechanism of BC and the development of drugs and diagnosis and treatment methods in the FISH analysis of ERBB2 and its related genes.
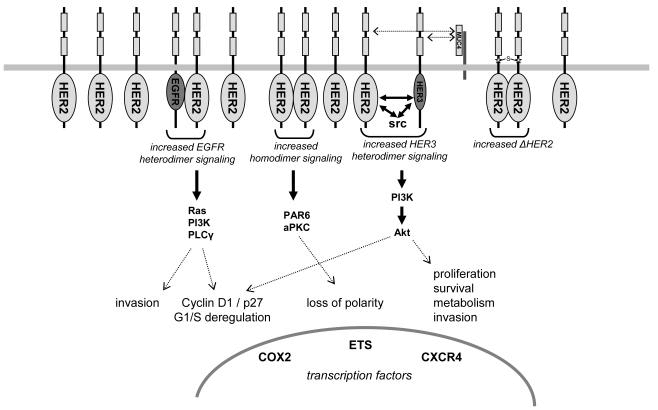 Fig 1. Schematic of the signaling abnormalities resulting from HER2 overexpression that are felt to contribute to tumorigenesis. (Moasser M M, et al. 2007)
Fig 1. Schematic of the signaling abnormalities resulting from HER2 overexpression that are felt to contribute to tumorigenesis. (Moasser M M, et al. 2007)
Analysis Service Content
The latest ASCO-CAP guidelines for HER2 detection for breast cancer have changed the assessment criteria for HER2 amplification. Specifically, it means that the average number of HER2 genes per tumor cell and the ratio of the average HER2 to the internal control chromosome 17 centromere need to be formally evaluated to assess HER2 status through FISH. We provide FISH analysis for ERBB2 and other genes in breast cancer samples, and we also provide IHC services for these markers to help researchers evaluate the ERBB2, ER, PR and other biomarkers in the sample. DNA-FISH and RNA-FISH can be used to analyze the copy number of target genes in tissue samples. Our RNA-FISH can analyze related lncRNA and other non-coding sequences through highly specific probes. In addition, our IHC service can evaluate the protein level of the target of interest in the sample and comprehensively analyze the changes in the target gene. The basic process of the service is program communication, sample mailing, sample analysis, data collection, data analysis and report submission. Customers can choose one of the analysis or combination analysis services. Our data analysis includes scoring HER2 protein expression and gene copy number scores based on the latest ASCO/CAP guidelines.
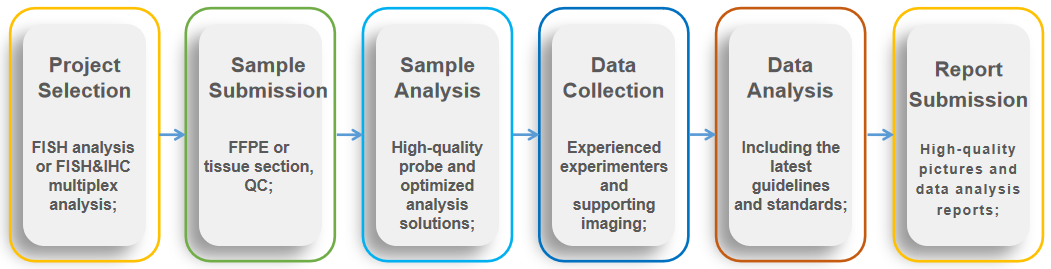 Fig 2. Service flow of in situ analysis available in breast cancer.
Fig 2. Service flow of in situ analysis available in breast cancer.
Creative Bioarray provides FISH and IHC technical services for breast cancer related research. Whether it is copy number, structural variation, mRNA analysis, ncRNA analysis or protein level evaluation. Our experimental platform has experienced molecular biology experts to escort your research.
If you are interested in our services, please contact us for cooperation. We look forward to cooperating with you in the near future.
References
- Moasser M M. The oncogene HER2: its signaling and transforming functions and its role in human cancer pathogenesis[J]. Oncogene, 2007, 26(45): 6469-6487.
- Press M F, Seoane J A, Curtis C, et al. Assessment of ERBB2/HER2 status in HER2-equivocal breast cancers by FISH and 2013/2014 ASCO-CAP guidelines[J]. JAMA oncology, 2019, 5(3): 366-375.


 Fig 1. Schematic of the signaling abnormalities resulting from HER2 overexpression that are felt to contribute to tumorigenesis. (Moasser M M, et al. 2007)
Fig 1. Schematic of the signaling abnormalities resulting from HER2 overexpression that are felt to contribute to tumorigenesis. (Moasser M M, et al. 2007) Fig 2. Service flow of in situ analysis available in breast cancer.
Fig 2. Service flow of in situ analysis available in breast cancer.


