In situ hybridization, especially its fluorescence-based variants (FISH), is commonly used in a wide range of disciplines, including genetics, developmental biology, pathology, and cell biology, to study the genome composition or gene expression in cells. Since the introduction of in situ hybridization technology in 1969, related technologies have been improved with changes in analysis requirements in terms of probes, labels, hybridization schemes, and microscope systems. FISH can help researchers understand the organization and composition of the genome and its aberrations, as well as the expression and location of gene transcripts in a visual way. However, the development of the FISH method is mainly used to detect fixed, morphologically preserved specimens, and therefore provides a static rather than dynamic view of nucleic acid positioning. The development of live cell imaging technology has improved this situation. In order to detect and track specific endogenous RNA in living cells, techniques based on in vivo hybridization have been developed. Using a variety of different nucleic acid probe types and fluorescence detection methods, methods for visualizing intracellular sequences of living cells have been gradually improved. Molecular beacons (MB) are stem-loop hairpin oligonucleotides with a fluorescent quencher at one end and a fluorescent dye (also called reporter gene or fluorophore) at the other end. This structure allows MB to not fluoresce in the absence of its target complementary sequence. Due to the spatial separation of the quencher and reporter molecule, MB will fluoresce after binding to the target. Molecular beacons are promising probes for the development of RNA imaging technology.
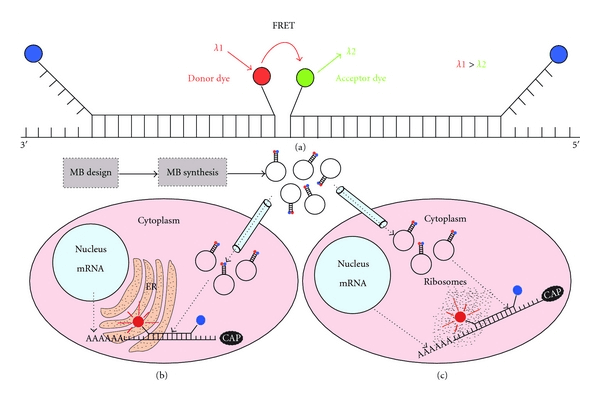 Fig 1. Approaches for RNA visualization in living cells. (Monroy-Contreras R, et al. 2011)
Fig 1. Approaches for RNA visualization in living cells. (Monroy-Contreras R, et al. 2011)
Transcriptomics Analysis Based on Molecular Beacons and FISH
Using the cell's own replication or transcription mechanism or by combining DNA or RNA-related fluorescent dyes, DNA or RNA molecules can be observed in living cells. Fluorescent in vivo hybridization is the preferred method for detecting specific endogenous RNA species in living cells. This service aims to use FISH and molecular beacons for transcriptomics analysis. Biological events involving coding RNA and non-coding RNA in cells will affect cell functions. Combining fluorescently labeled molecular beacons with fluorescence detection can realize RNA imaging before FISH analysis and obtain information on RNA expression in living cells.
This service is mainly carried out for cell samples, using customized Peptide Nucleic Acid Molecular Beacons to be incorporated into cells, using confocal imaging equipment to observe and harvest data images. At the same time, we also provide analysis based on flow cytometry. The cells were then fixed and analyzed by RNA-FISH. Through the integration of live cell fluorescence imaging data and FISH data, expand the field of view and gain more knowledge. The specific PNA molecular beacon has better targeting than DNA probes and a low mismatch rate. Our expert team will help customers complete the sequence design of molecular beacons, including the selection of dyes and quenchers.
Sample type: The submitted cell sample needs to be no less than 105;
Target molecules: transcripts (mRNA) and target RNA molecules (sRNA, microRNA);
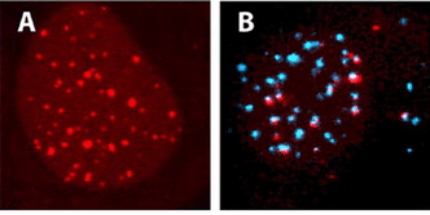 Fig 2. Visualization of nucleic acids in living osteosarcoma U2OS cells by fluorescence confocal microscopy. (Dirks R W, et al. 2006)
Fig 2. Visualization of nucleic acids in living osteosarcoma U2OS cells by fluorescence confocal microscopy. (Dirks R W, et al. 2006)
Creative Bioarray uses PNA molecular beacon probes to implement live cell RNA tracking and FISH hybridization services, helping our customers achieve the dual analysis requirements of live cell RNA tracking and FISH positioning. You will benefit from our technical expertise and complete platform, and work with you to find the best solution to meet your needs. If you have live cell RNA tracking and RNA-FISH analysis needs or are interested in our services, please contact us for cooperation. We look forward to cooperating with you in the near future.
References
- Monroy-Contreras R, Vaca L. Molecular beacons: powerful tools for imaging RNA in living cells[J]. Journal of nucleic acids, 2011, 2011.
- Dirks R W, Tanke H J. Advances in fluorescent tracking of nucleic acids in living cell[J]. Biotechniques, 2006, 40(4): 489-496.





