Fiber-FISH
Fiber fluorescence in situ hybridization (Fiber-FISH) is a high-resolution FISH technology active in the 1990s. It allows high-resolution mapping of genes and chromosomal regions on chromatin or DNA fibers. At the same time, this technique enables the physical sequencing of DNA probes to a resolution as low as 1000 bp, and also allows the assessment of gaps and overlaps in contigs, as well as analysis of segment duplication and copy number variation. In practice, chromatin/DNA fibers are released from the cell nucleus, usually by salt or solvent extraction, and they are stretched and fixed on a microscope slide before hybridization. Fiber-FISH provides higher resolution than ordinary FISH and produces more accurate information about the location of specific DNA probes on chromosomes. Fiber-FISH analysis of meiotic (pachytene) chromosomes is also called Pachytene-FISH. It has been reported that high-resolution thick-line phase FISH and fiber FISH have been used to determine the location of some "chirality gene". In addition, Fiber-FISH can be combined with other technologies to expand the application range, such as single-cell genetic characterization by integrating chromosomal DNA microfluidic stretching and fiber-FISH.
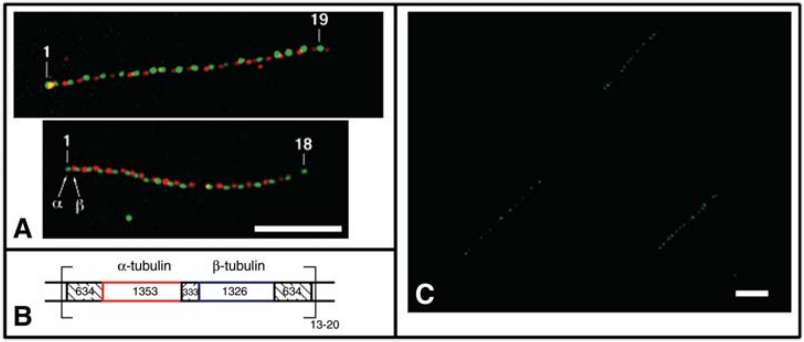 Fig 1. Detection of repetitive gene organization by Fiber-FISH. (Ersfeld K et al., 1994)
Fig 1. Detection of repetitive gene organization by Fiber-FISH. (Ersfeld K et al., 1994)
Technical Features
Fiber-FISH can be used for physical mapping and detection of genomic rearrangements in the 1-300 kb range. It hybridizes to immobilized linearized DNA using region-specific probes.
The conventional Fiber-FISH method is to prepare extended chromatin fibers from chromosomes in living cells, a process of preparing chromatin fibers by in situ treatment of cells attached to a glass slide. This method uses detergents or high-concentration salt extraction to deproteinize chromatin. This treatment completely unwinds the DNA to a linear duplex level of approximately 340 nm/1000 bp. Fiber slides prepared after protein removal were aged overnight in a sealed box with desiccant and hybridized the next day. In addition, this technique can also be performed on the preparation of "naked DNA" in protease-treated cells, i.e., chromosome separation after pulsed-field gel electrophoresis (PFGE). This treatment causes the DNA to split into fragments of different sizes. Fiber slides prepared by the above procedure were aged overnight and used for FISH hybridization.
The Pachytene-FISH method is for meiotic (pachytene) chromosomes, and the preparation of fiber slides is the same as above.
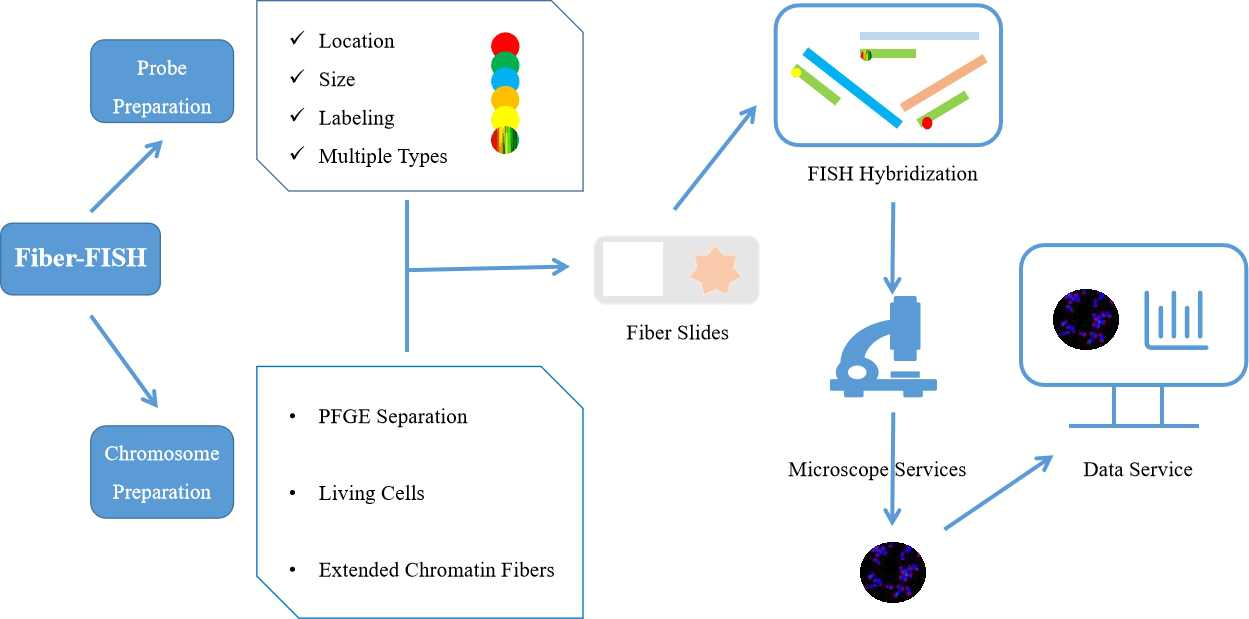 Fig 2. Comprehensive fiber-FISH services.
Fig 2. Comprehensive fiber-FISH services.
Creative Bioarray provides high-resolution FISH testing services to help researchers access chromosomes. The early exploration of Fiber-FISH technology promoted the improvement of the resolution of FISH technology, which further expanded the application scope of FISH technology. By choosing our FISH technology you will benefit from our technical expertise and high-resolution FISH experimental platform. If you are interested in our FISH service, please contact us for cooperation. We look forward to working with you in the near future and working with you to find the best solution to meet your needs.
References
- Ersfeld K. Fiber-FISH: fluorescence in situ hybridization on stretched DNA[M]//Parasite Genomics Protocols. Humana Press, 1994: 395-402.
- Liu M M, Davey J W, Banerjee R, et al. Fine mapping of the pond snail left-right asymmetry (chirality) locus using RAD-Seq and fibre-FISH[J]. PloS one, 2013, 8(8): e71067.
All products and services on this website are only suitable for non-medical purposes.


 Fig 1. Detection of repetitive gene organization by Fiber-FISH. (Ersfeld K et al., 1994)
Fig 1. Detection of repetitive gene organization by Fiber-FISH. (Ersfeld K et al., 1994) Fig 2. Comprehensive fiber-FISH services.
Fig 2. Comprehensive fiber-FISH services.


