Methyl-specific FISH
Methyl-specific FISH Technology
DNA methylation is an important epigenetic modification of many plant and animal genomes, such as 5-methylcytosine (5mC), which mainly occurs on cytosine bases. DNA methylation and demethylation significantly affect the inactivation and activation processes of gene expression, respectively. Determining the location and frequency of DNA methylation is important for elucidating the molecular mechanisms of cell differentiation and some disease states. Methylation-specific fluorescence in situ hybridization (MeFISH) was developed to microscopically visualize the DNA methylation status of specific repetitive sequences in a single cell. This technology is based on the different reactivity of 5-methylcytosine and cytosine in the target DNA to form a sequence-specific osmium complex.
Sequence-specific osmium complexation is achieved by hybridizing short DNA molecules containing the functional nucleotides to the target DNA sequence, resulting in the formation of cross-linked structures. The huge difference in the oxidation rate of osmium at modified and unmodified sites allows the interstrand complex formed by osmium and nucleic acid (ICON) probes to allow sequence-selective detection of 5mC in vitro. The method of using synthetic fluorescent-labeled ICON probes to hybridize with cell nuclei and chromosomes can be used to analyze the methylation status of major and minor satellite repeats in experimental animals.
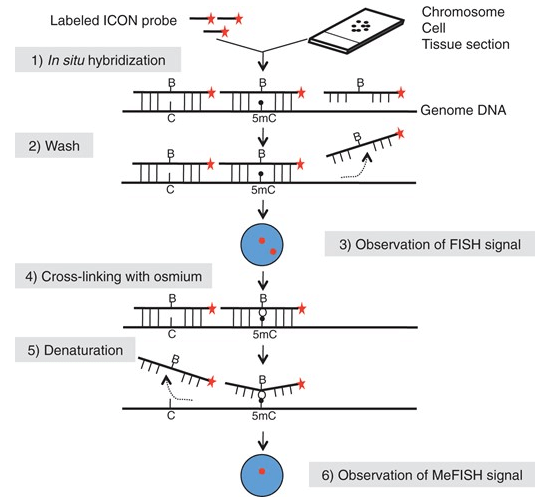 Fig 1. Outline of the MeFISH protocol. (Li Y et al., 2013)
Fig 1. Outline of the MeFISH protocol. (Li Y et al., 2013)
MeFISH-based Methylation Analysis
MeFISH technology can visualize the DNA methylation status of specific target sequences in single-cell nuclei or chromosomes. The technology can be performed on cell or tissue section samples, which researchers can test with custom-made ICON probes and assess the methylation status in the samples. The detection principle of MeFISH is to achieve sequence-specific osmium complexation by hybridizing short DNA molecules containing functional nucleotides with target DNA sequences to form cross-linked structures. The general steps of MeFISH include section pretreatment, probe preparation and hybridization, non-specific binding removal, osmium treatment, non-cross-linked probe removal, and signal collection. Interstrand crosslinks clearly distinguish methylated and unmethylated cytosines and are used to quantify the degree of methylation of specific cytosines in the genome. In addition, researchers also need to analyze 5hmC, which can be solved by a combination of immunostaining and MeFISH. Our FISH technology platform provides customization of ICON probes.
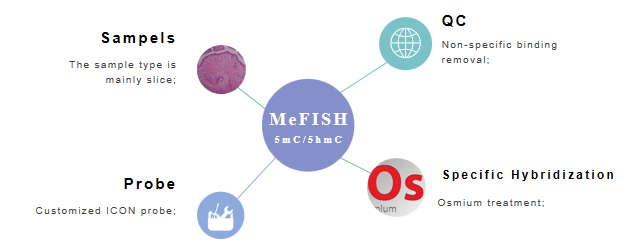 Fig 2. The main points of MeFISH to detect methylation.
Fig 2. The main points of MeFISH to detect methylation.
Technology Features
- Detect 5mC and 5hmC in the sample
- In-situ visualization method of methylation
- Combined immunostaining to analyze 5mC/5hmC
Creative Bioarray provides services based on custom ICON probes to assist researchers in the analysis of methylation through smoke-flavor technology. You will benefit from our technology platform and professional service team. If you are interested in this service, please contact us for cooperation. We look forward to cooperating with you in the near future.
References
- Li Y, Miyanari Y, Shirane K, et al. Sequence-specific microscopic visualization of DNA methylation status at satellite repeats in individual cell nuclei and chromosomes[J]. Nucleic acids research, 2013, 41(19): e186-e186.
- Shiura H, Okamoto A, Sasaki H, et al. Whole-mount MeFISH: a novel technique for simultaneous visualization of specific DNA methylation and protein/RNA expression[J]. PloS one, 2014, 9(4): e95750.
All products and services on this website are only suitable for non-medical purposes.


 Fig 1. Outline of the MeFISH protocol. (Li Y et al., 2013)
Fig 1. Outline of the MeFISH protocol. (Li Y et al., 2013) Fig 2. The main points of MeFISH to detect methylation.
Fig 2. The main points of MeFISH to detect methylation.


