ReD-FISH and Telomere Replication
ReD-FISH Technology
Telomeres are located at the ends of all eukaryotic chromosomes and are essential for ensuring chromosomal integrity and maintaining overall genome stability. Vertebrate chromosome ends generally consist of telomeric repeats (a series of identical tandem TTAGGG sequences). These sequences bind to histone proteins to form specialized structures to protect chromosomes from end-to-end fusion, degradation, and inappropriate recombination. Telomeres are essential for the complete replication of eukaryotic chromosomes, but telomere replication in mammalian cells is associated with replication throughout the S phase. Based on the chromosome-directed-FISH (CO-FISH) procedure, replication-de-targeted FISH (ReD-FISH) was developed as a unique tool to study telomere replication patterns located on individual chromosome arms. This method is also suitable for examining telomeres in species belonging to different animal classes, for which well-proliferating cell cultures can be established and maintained. ReD-FISH can be used to study telomere replication patterns at each chromosome end at defined stages of the S phase. Recently, highly specific peptide nucleic acid (PNA) and locked nucleic acid (LNA) telomeric probes have been reported to replace DNA probes as nucleic acid analogs for ReD-FISH analysis. Such probes incorporating nucleotide analogs have better binding specificities than DNA probes.
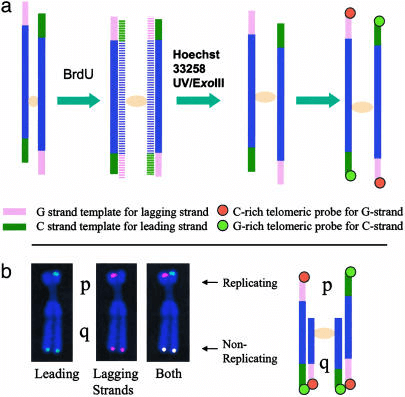 Fig 1. Schema of ReDFISH technique. (Zou Y et al., 2004)
Fig 1. Schema of ReDFISH technique. (Zou Y et al., 2004)
ReD-FISH Assays in Telomere Replication Research
ReD-FISH experiments can be performed using one of two brominated analogs, BrdU or BrdC, in this way the timing of replication of specific sequences can be determined. The principle is simply that if BrdU is incorporated into the sequence of interest due to replication, the newly formed DNA strand will be detargeting. Each oligonucleotide probe will hybridize to only one of the parental strands, and only one chromatid will show a signal. However, if the sequence of interest has not been replicated and BrdU has not been incorporated into the newly synthesized strand, a standard double signal will be displayed on both chromatids of the metaphase chromosome. In experiments for telomere replication studies, if telomeres were fully replicated during BrdU treatment, the ReD-FISH signal was only present on one of the two sister chromatids. If telomeres replicate in another time window, the ReD-FISH signal will be visible on both sister chromatids.
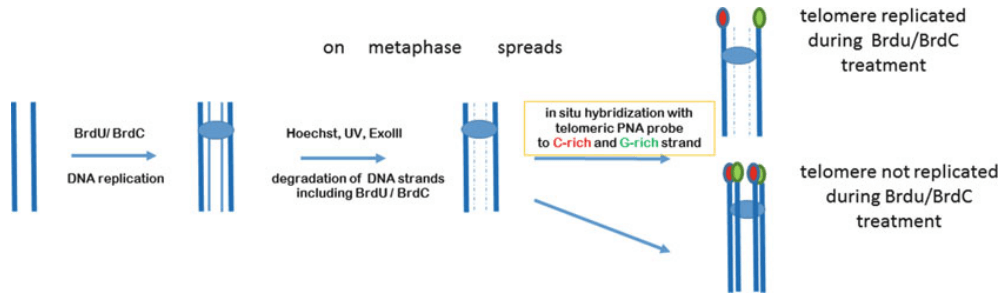 Fig 2. A ReD-FISH protocol to study telomere replication. (Rubtsov N et al., 2017)
Fig 2. A ReD-FISH protocol to study telomere replication. (Rubtsov N et al., 2017)
Applications of ReD-FISH
- ReD-FISH provides qualitative and quantitative information on telomere replication timing
- The knowledge gained by ReD-FISH can be used to analyze the relationship between telomere replication timing defects and chromosome condensation defects
- The knowledge gained by ReD-FISH can be used to analyze sister chromatid cohesion
- The knowledge gained by ReD-FISH can be used to analyze genome stability
Creative Bioarray provides high-quality probe products and probe customization services that can apply a variety of FISH technology variants and methods. If you are interested in our service, please contact us for cooperation. We look forward to cooperating with you in the near future.
References
- Zou Y, Gryaznov S M, Shay J W, et al. Asynchronous replication timing of telomeres at opposite arms of mammalian chromosomes[J]. Proceedings of the National Academy of Sciences, 2004, 101(35): 12928-12933.
- Rubtsov N, Zhdanova N. The replicative detargeting FISH (ReD-FISH) technique in studies of telomere replication[M]//Fluorescence In Situ Hybridization (FISH). Springer, Berlin, Heidelberg, 2017: 159-168.
All products and services on this website are only suitable for non-medical purposes.


 Fig 1. Schema of ReDFISH technique. (Zou Y et al., 2004)
Fig 1. Schema of ReDFISH technique. (Zou Y et al., 2004) Fig 2. A ReD-FISH protocol to study telomere replication. (Rubtsov N et al., 2017)
Fig 2. A ReD-FISH protocol to study telomere replication. (Rubtsov N et al., 2017)


